data anlysis made easy
Muscle-BIDS step by step
To use automated processing of muscle bids data QMRITools follows three main steps:
- BIDS converstion
- Processing
- Analysis
The challenge behind automated processing is actually understading the data and knowing how to handle it. Here there are many choices and not all can be drived from the data itself. The approach in QMRITools to handle this is by specifying these choises in a config.json file. Of which an example is shown a the bottom of this page.
Starting a project
To start processing the raw dicom data should be placed in the source folder where each subject en session is its own folder and where the folder name is comprised of this subject name and session code.
Subject001
Subject002_001
Subject002_002
etc..
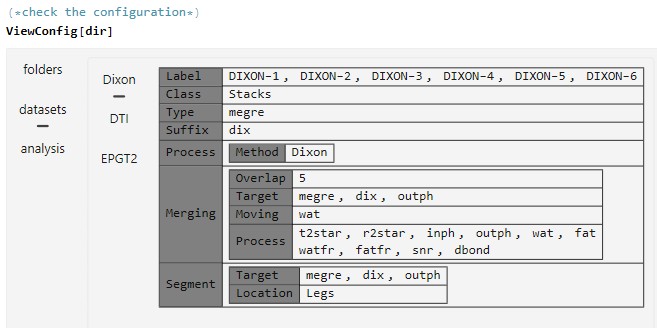
To start processing the configuration file is placed in the same directory dir which is given to the software as input. A configured config.json file can be checked and evaluated by the commands.
ViewConig[dir]
CheckConfigLabels[dir]
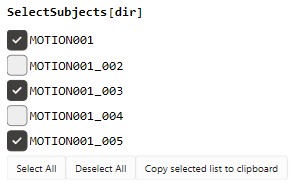
If a subset of data needs to be processed this can be done with
SelectSubjects[dir]
Step 1. Bids conversion
(1. convert the dcm to nii - Skips the folders already done)
BidsDcmToNii[dir]
ViewProtocolNames[dir, ProcessSubjects -> subs]
(2. convert the raw nii to bids nii, deletes raw files after conversion)
MuscleBidsConvert[dir]
Step 2. Data processing
(3. process all bids files that are found, processing is in config)
MuscleBidsProcess[dir, ProcessSubjects -> subs, VersionCheck -> False]
(4. merge multiple stacks of the same data into one dataset)
MuscleBidsMerge[dir, ProcessSubjects -> subs, VersionCheck -> False]
(4. perform the segmentation using CNN)
MuscleBidsSegment[dir, ProcessSubjects -> subs, VersionCheck -> False]
(6. Perform tractography, generates tracts)
MuscleBidsTractography[dir, ProcessSubjects -> subs, VersionCheck -> False]
Step 2. Analysis of output
(7. Perform data analsyis, generates xls and images) MuscleBidsAnalysis[dir, ProcessSubjects -> subs]
Example of config.json
{
"folders":{
"dicomData":"01_sourcedata",
"rawData":"02_rawdata",
"derivedData":"03_derivatives",
"mergeData":"04_merged",
"analysis":"05_analysis"
},
"conversion":{
"Version": "23"
},
"datasets":{
"Dixon":{
"Label":["GE_Dix_1", "GE_Dix_2"],
"Class":"Stacks",
"Type":"megre",
"Suffix":"dix",
"Process":{
"Method":"Dixon"
},
"Merging":{
"Overlap":5,
"Target":["megre", "dix", "wat"],
"Moving":"wat",
"Process":[ "inph", "outph", "wat", "fat", "fatfr"]
},
"Segment":{
"Target":["megre", "dix", "outph"],
"Location":"Legs",
"Device":"CPU"
}
},
"T2map":{
"Label":["MESE_15echo_1", "MESE_15echo_1"],
"Class":"Stacks",
"Type":"mese",
"Suffix":"t2",
"Process":{
"Method":"EPGT2",
"Settings":[
["exitation", 90, [1.5991, 5.661, 538]],
["echo", 180, [1.3326, 2.9376, 340]]
],
},
"Merging":{
"Overlap":[5, 3],
"Target":["megre", "dix", "wat"],
"Moving":"wat",
"Process":["t2w", "t2f", "wat", "fat", "fatfr"]
}
},
"DTI":{
"Label":["DTI_30dir_1","DTI_30dir_2"],
"Class":"Stacks",
"Type":"dwi",
"Suffix":"dti",
"Process":{
"Method":"DTI",
"FlipPermute": [[1, -1, 1], ["z", "y", "x"]],
"GradientCorrection": true
},
"Merging":{
"Overlap":5,
"Target":["megre", "dix", "outph"],
"Moving":"s0",
"Process":["s0", "snr0", "l1", "l2", "l3", "md",
"fa", "rd", "adci", "fri", "tens"]
},
"Tractography":{
"Target":["dwi", "dti", "tens"],
"Segmentation":["seg", "auto", "megre", "dix", "outph"],
"Stopping":[
[["dwi", "dti", "fa"], [0.05, 0.65]],
[["dwi", "dti", "md"], [0.5, 2.5]]
],
"FlipPermute":[[1, -1, 1], ["z", "y", "x"]]
}
}
},
"analysis":{
"Segmentation":{
"Type":["seg", "auto", "megre", "dix", "outph"],
"Labels":[90, "Legs"]
},
"Analysis":{
"Types":[
["dwi", "dti", ["l1", "l2", "l3", "md", "rd", "fa", "adci", "fri", "snr0"]],
["dwi", "dti", "trk", ["leng", "ang", "seed", "dens"]],
["megre", "dix", ["fatfr", "t2star"]],
["mese", "t2", ["fatfr", "t2w", "t2f"]]
],
"TractBased":[
["dwi", "dti", ["l1", "l2", "l3", "md", "rd", "fa", "adci", "fri", "snr0"]],
["dwi", "dti", "trk", ["leng", "ang"]]
],
"Options":{
"MaskErosion":true,
"TractWeigthing":false
}
},
"Images":{
"Reference": ["megre", "dix", "wat"],
"TractImages": ["dwi", "dti", "trk", "plot"],
"QuantImages":[
[["megre", "dix", "fatfr"], "Lajolla", [0.001, 0.999], ["0", "100", "Fat Frac. [%]"]],
[["megre", "dix", "t2star"], "Navia", [0.0001, 0.1], ["0", "100", "T2* [ms]"]],
[["megre", "dix", "outph"]],
[["dwi", "dti", "fa"], "Devon", [0.001, 0.75], ["0", "1", "FA [-]"]],
[["dwi", "dti", "md"], "Lapaz", [0.5, 2.0], ["0.5", "2", "MD 10^-3 mm^2/s"]],
[["dwi", "dti", "rd"], "Lapaz", [0.5, 2.0], ["0.5", "2", "RD 10^-3 mm^2/s"]],
[["dwi", "dti", "snr0"], "Batlow", [1, 50], ["0", "50", "SNR [-]"]],
[["dwi", "dti", "s0"]],
[["dwi", "dti", "trk", "seed"], "BlackToWhite", [0, 2]],
[["dwi", "dti", "trk", "dens"], "CherryTones", [0.01, 5], ["0", "5", "Tract dens. [-]"]],
[["dwi", "dti", "trk", "leng"], "Roma", [1, 250], ["0", "250", "Tract length [mm]"]],
[["dwi", "dti", "trk", "curv"], "Bilbao", [0.1, 30], ["0", "30", "Tract curvature [1/m]"]],
[["dwi", "dti", "trk", "ang"], "Vik", [0.1, 90], ["0", "90", "Tract angle [Degree]"]],
[["mese", "t2", "t2w"], "Navia", [5, 45], ["5", "45", "T2 water [ms]"]],
[["mese", "t2", "wat"]]
]
}
}
}